Physics-informed convolutional-recurrent neural networks for solving spatiotemporal PDEs.
Paper link: [Journal Paper], [ArXiv]
By: Pu Ren, Chengping Rao, Yang Liu, Jian-Xun Wang and Hao Sun
This repository is maintained by the authors, with assistance from GPT-5.4.
PhyCRNet is a physics-informed convolutional recurrent architecture for learning spatiotemporal dynamics of partial differential equations. This repository contains:
- high-order finite-difference solvers for dataset generation,
- the Burgers-equation PhyCRNet model implementation,
- a training and evaluation entrypoint,
- utility functions for checkpointing, plotting, and experiment management.
The current codebase keeps the original research content while organizing the project into clearer modules for day-to-day development.
- Present a Physics-informed discrete learning framework for solving spatiotemporal PDEs without any labeled data
- Proposed an encoder-decoder convolutional-recurrent scheme for low-dimensional feature extraction
- Employ hard-encoding of initial and boundary conditions
- Incorporate autoregressive and residual connections to explicitly simulate the time marching
We show the comparison between PhyCRNet and PINN on 2D Burgers' equations below. The left, middle and right figures are the ground truth, the result from our PhyCRNet and the result from PINNs respectively.
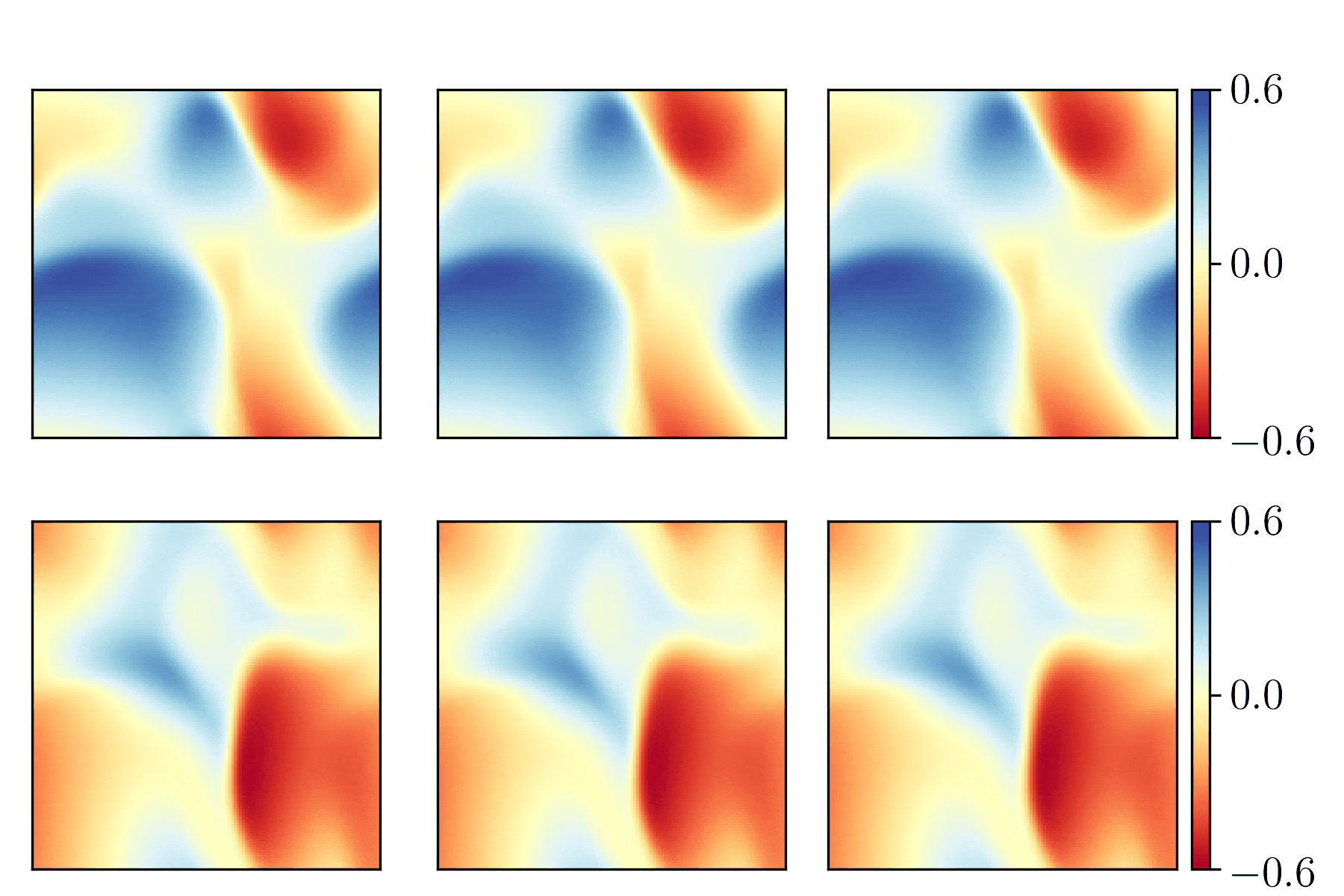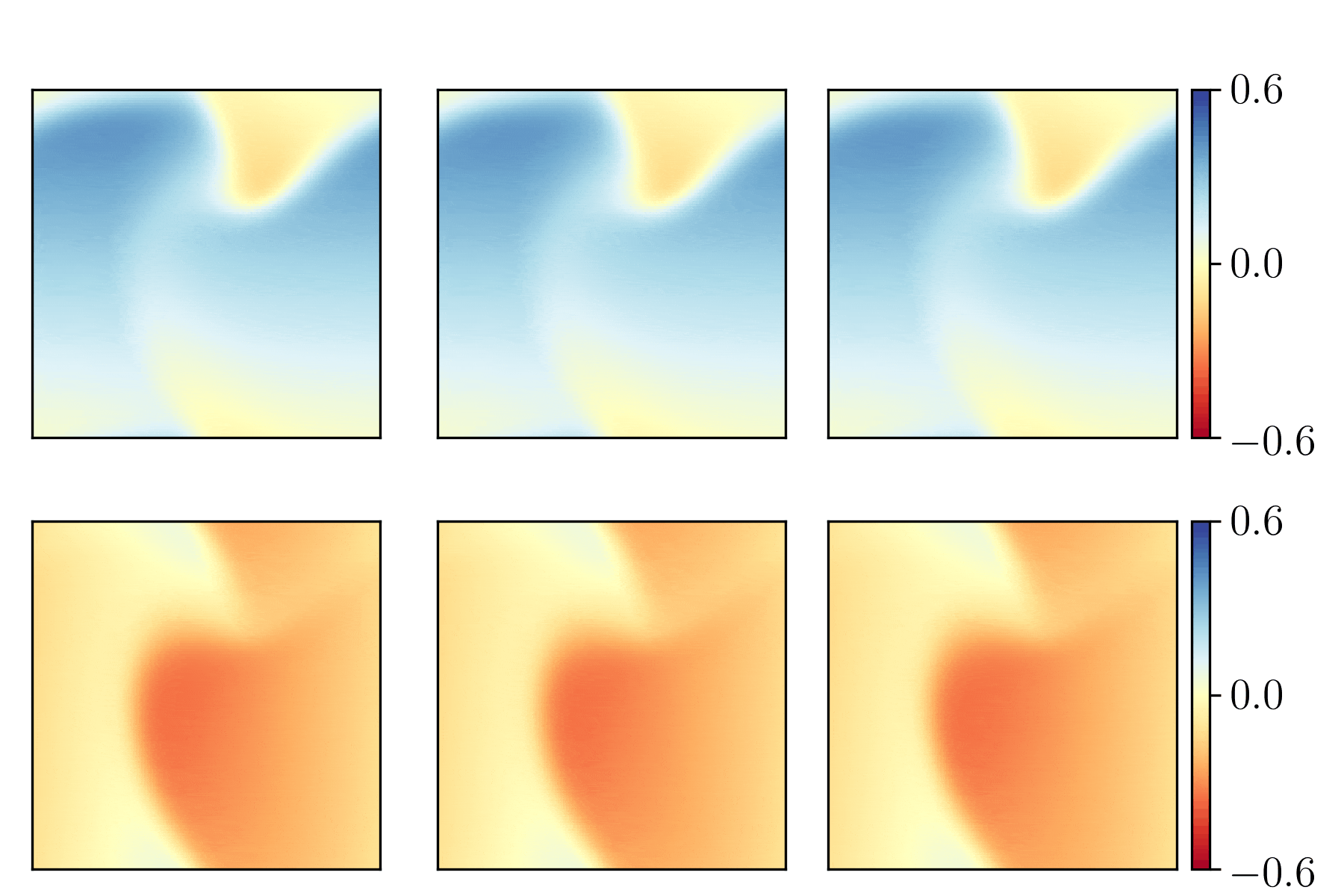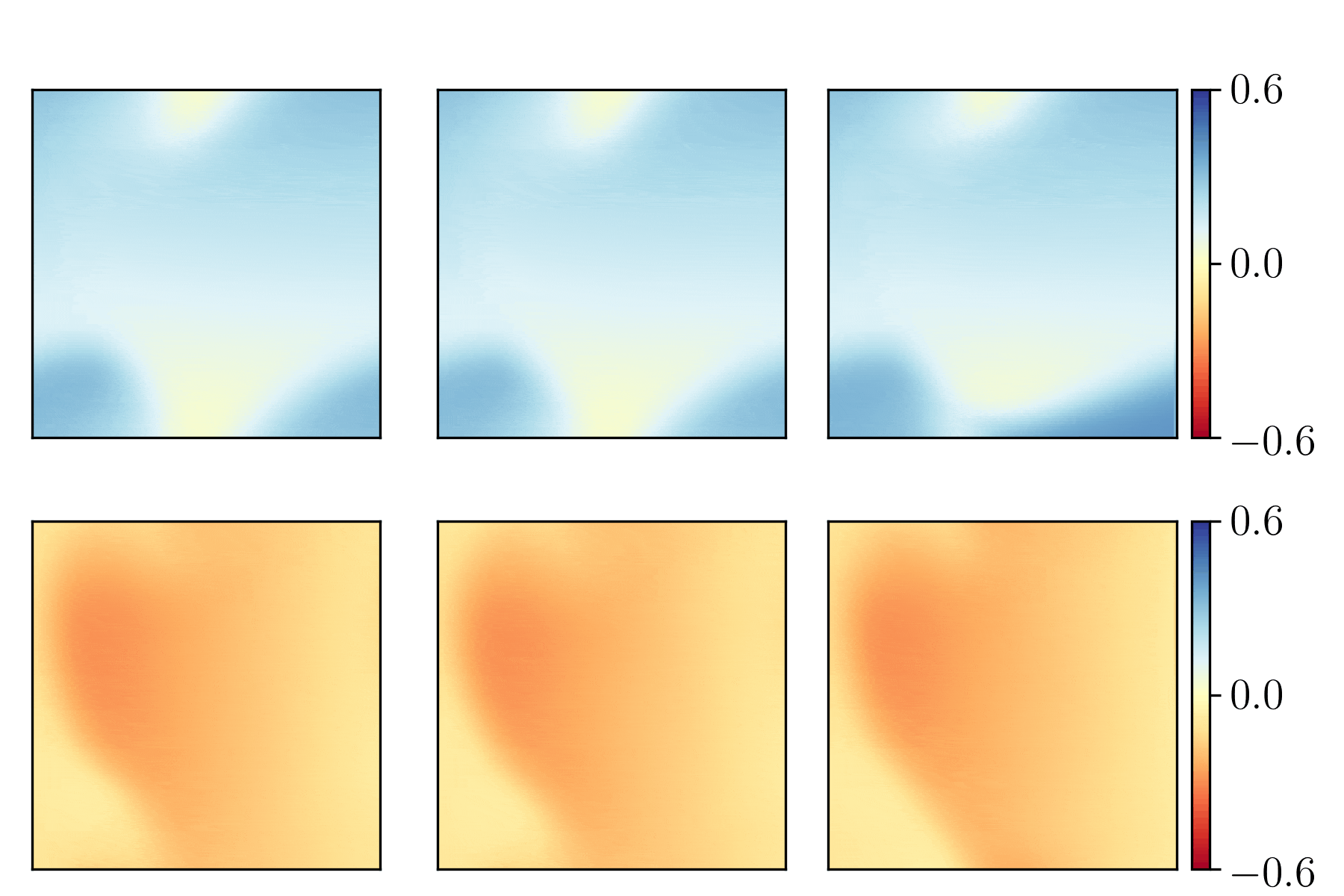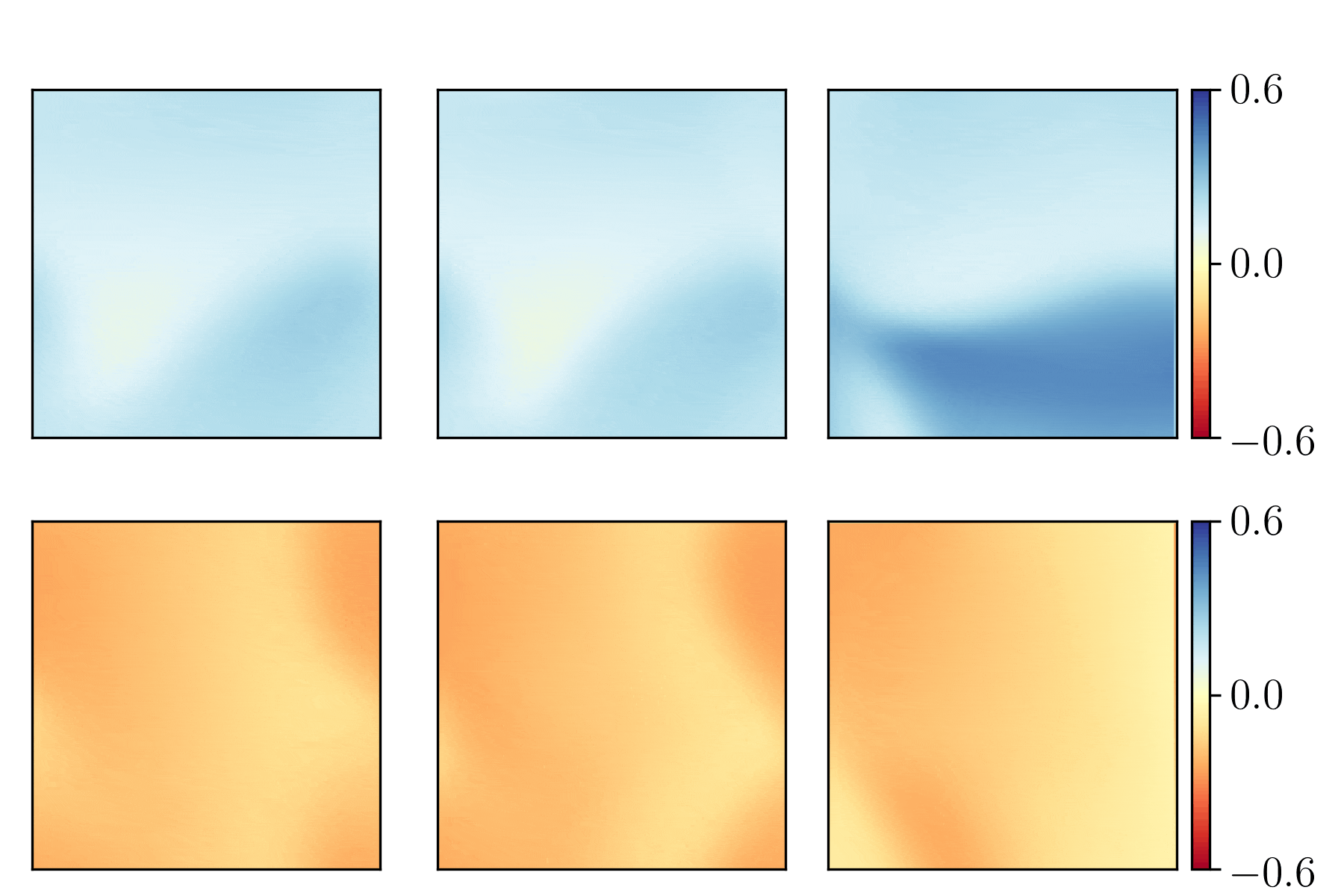
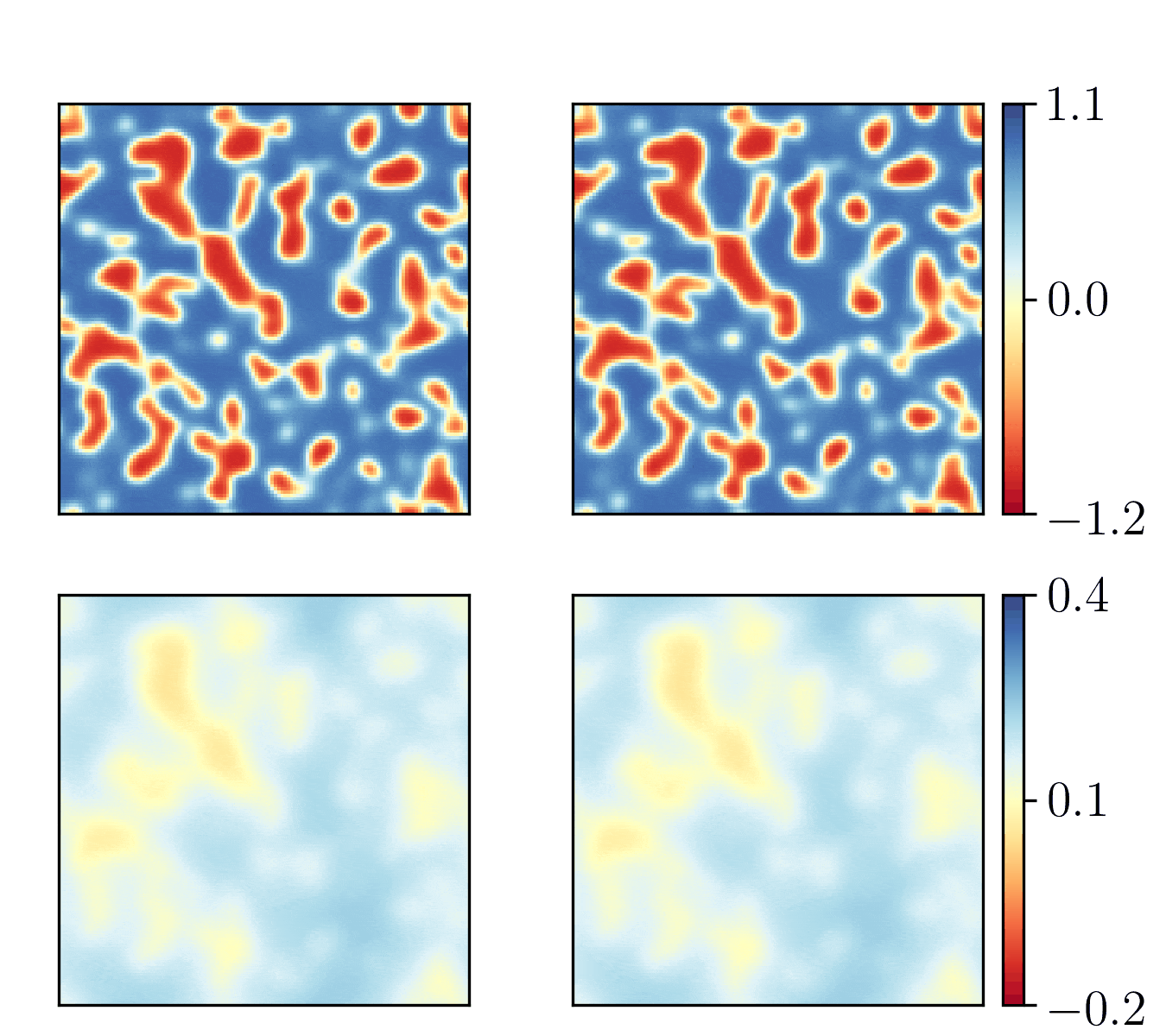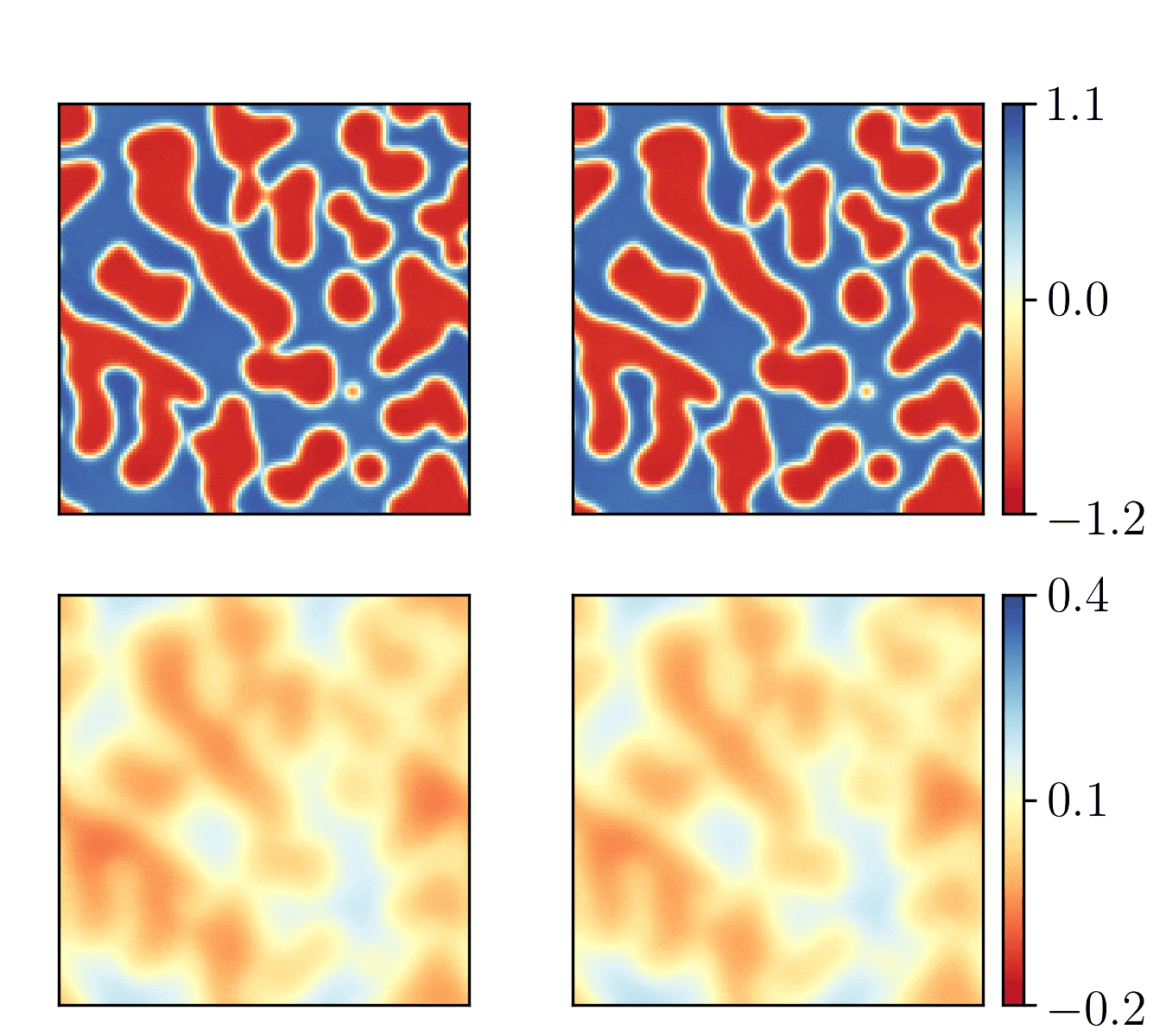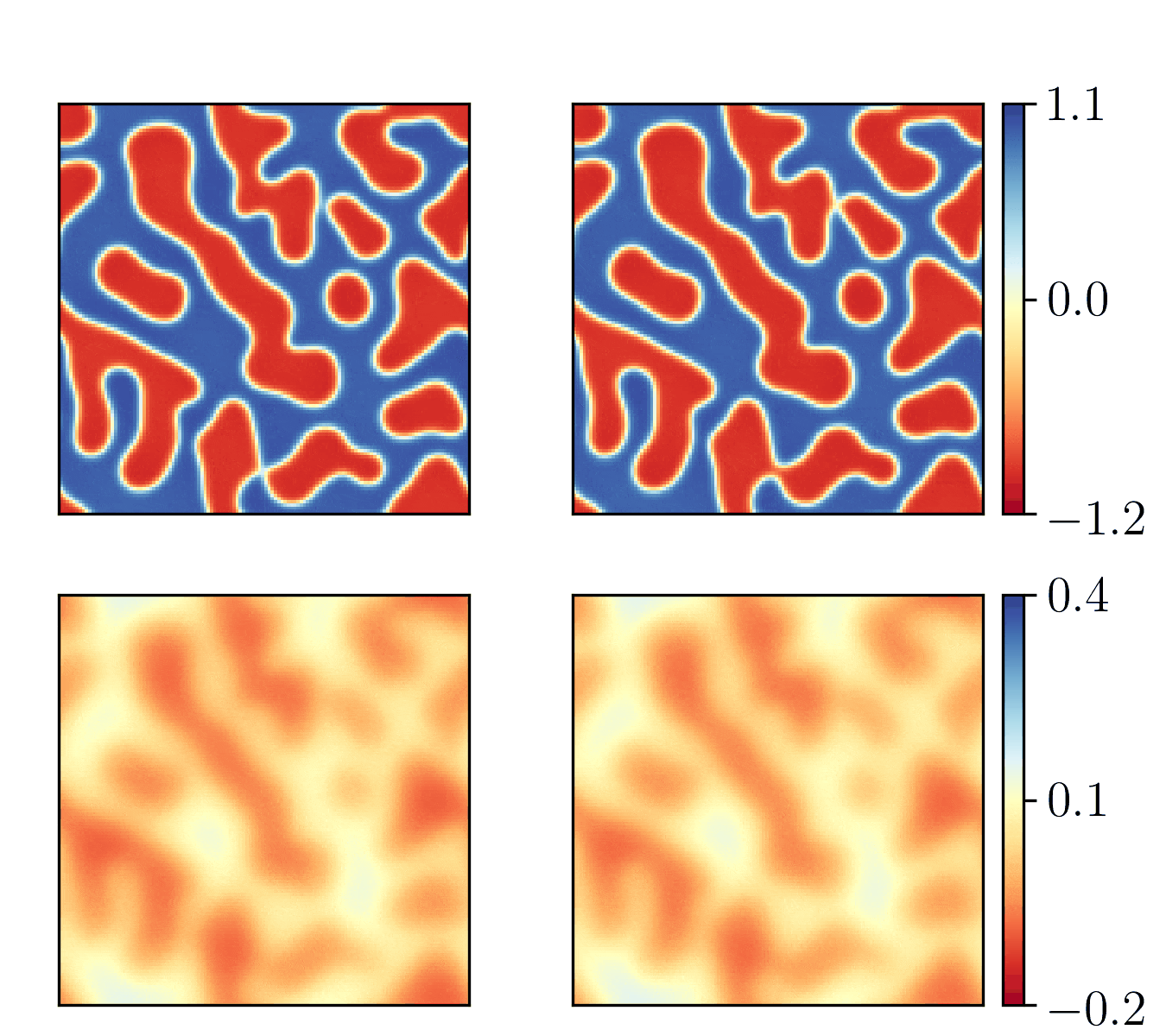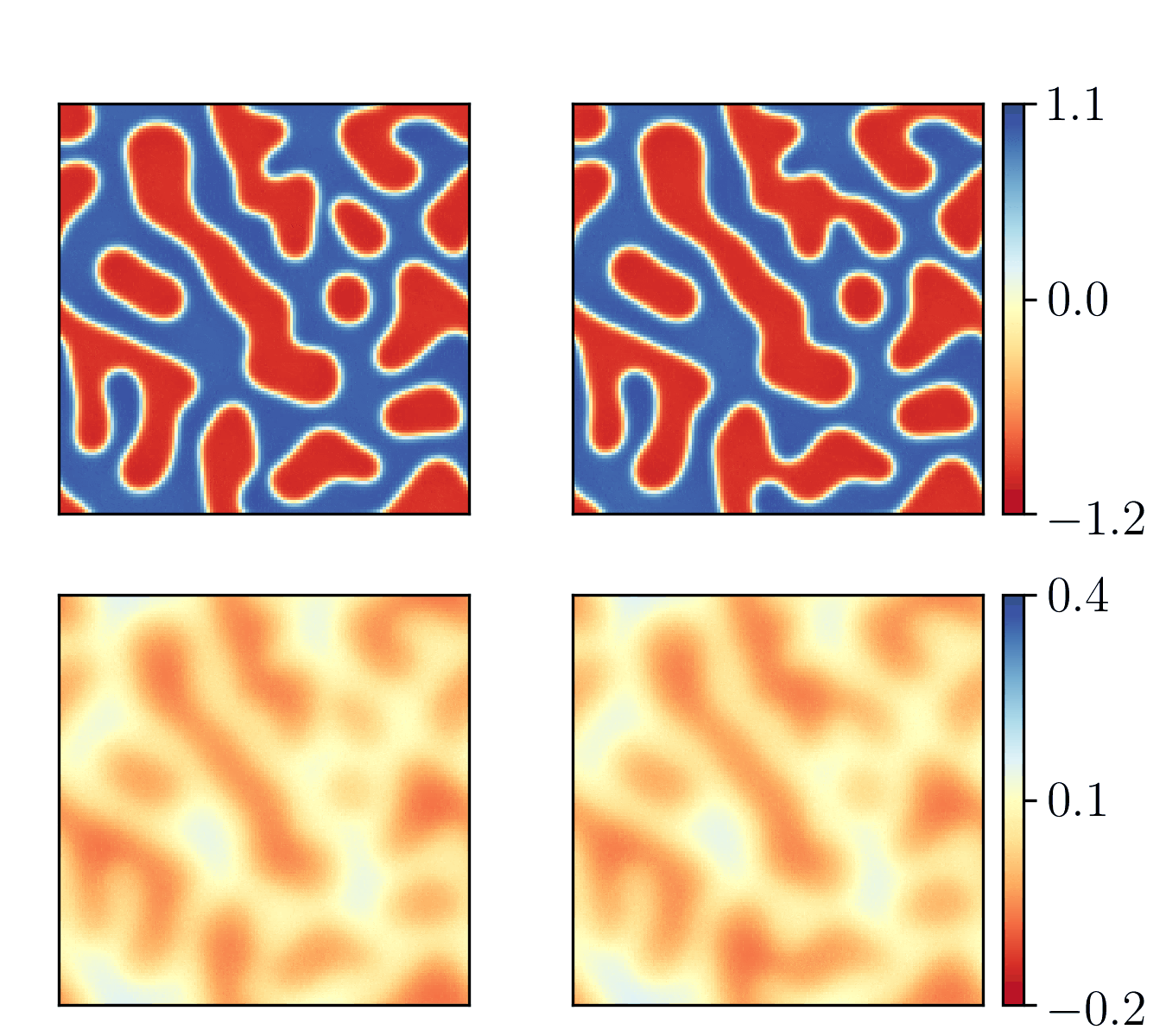
We show the generalization test on FitzHugh-Nagumo reaction-diffusion equations with four different initial conditions. The left and right parts are the ground truth generated with the high-order finite difference method and the results from our PhyCRNet, respectively.
PhyCRNet/
├── Datasets/
│ ├── Burgers_2d_solver_[HighOrder].py
│ ├── FN_2d_solver_[HighOrder].py
│ └── random_fields.py
├── Models/
│ └── PhyCRNet_burgers.py
├── train.py
├── utils.py
└── README.md
- Python 3.6.13
- Pytorch 1.6.0
- Other packages such as Matplotlib, Numpy and Scipy are also used
The repository does not currently ship with a pinned environment file, so installation is manual.
Example:
pip install numpy scipy matplotlib
pip install torch==1.6.0If you use a newer PyTorch version, parts of the original codebase may require additional compatibility updates.
We provide the codes for data generation used in this paper, including 2D Burgers' equations and 2D FitzHugh-Nagumo reaction-diffusion equations. They are coded in the high-order finite difference method. Besides, the code for random field is modified from [Link]. You may find the data solver for lambda-omega reaction-diffusion equations in [Link].
The initial conditions tested in this paper are also provided in the folder Datasets.
The solvers under Datasets/ can be used to generate trajectories for training or evaluation:
Datasets/Burgers_2d_solver_[HighOrder].pyDatasets/FN_2d_solver_[HighOrder].py
These scripts implement fourth-order finite-difference spatial discretization and RK4 time stepping.
The original project described the implementation as living under a general code folder. The current layout keeps that content but separates responsibilities more clearly:
Models/PhyCRNet_burgers.py: Burgers PhyCRNet architecture and physics-loss operatorstrain.py: training and evaluation entrypointutils.py: checkpointing, plotting, seeding, and helper utilities
The general code of PhyCRNet is provided in the repository using 2D Burgers' equations as a testing example. For other PDE systems, the network setting is similar. You may try modifying the grid sizes and time steps to your own cases.
It is important to pretrain the network from a small number of time steps (e.g., 100) and then gradually train on a longer dynamical evolution. Taking 2D Burgers' equation as an example, we pretrain the model from 100, then 200 and 500, and finally 1000. The model is able to extrapolate for at least another 1000 time steps.
python train.py --mode train
python train.py --mode eval
python train.py --mode allBy default, train.py expects:
- dataset at
./data/burgers_1501x2x128x128.mat - checkpoints under
./model/ - figures under
./figures/
- The repository is research-oriented rather than packaged as a library.
- File paths and training settings are currently configured directly in
train.py. - For reproducibility, random seeds are set in the data-generation and training code.
If you find our research helpful, please consider citing us with:
@article{ren2022phycrnet,
title={PhyCRNet: Physics-informed convolutional-recurrent network for solving spatiotemporal PDEs},
author={Ren, Pu and Rao, Chengping and Liu, Yang and Wang, Jian-Xun and Sun, Hao},
journal={Computer Methods in Applied Mechanics and Engineering},
volume={389},
pages={114399},
year={2022},
publisher={Elsevier}
}This project is distributed under the terms of the LICENSE file.